



Search websites, locations, and people

On the quest for the fluorescent “superheroes”
Kiryl Piatkevch Lab
26, 2024
Email: zhangchi@westlake.edu.cn
Phone: +86-(0)571-86886861
Office of Public Affairs
Prof. Kiryl Piatkevch's research team at Westlake University, in collaboration with Fedor Subach, reported a new extremely stable and bright green fluorescent protein, mBaoJin, which is suitable for super-resolution long-term live cell imaging and expansion microscopy.
Read the full story at https://www.nature.com/articles/s41592-024-02203-y

Since the 17th century, microscopy has driven many prominent breakthroughs and discoveries in life sciences, from the foundation of cell theory to mapping neuronal activity in the brain. Despite its 400-year history in biological research, microscopy still plays an increasingly essential role in new biological discoveries. Fluorescent proteins represent a crucial extension of light microscopy hardware as they enable real-time and multiplexed visualization of desired cellular structures and proteins through specific labeling on the genetic level. Furthermore, fluorescent probes can report various physiological activities ranging from membrane potential and redox status to small ions concentration and enzymatic activities. In this regard, basic and applied research in biology is becoming increasingly reliant on the advances in the engineering of novel fluorescent proteins that can extend the spatial and temporal scales of fluorescence imaging. There is no doubt that novel fluorescent probes that enable new types of biological measurements and optical recordings will help scientists deepen and broaden our understanding of the fundamentals of life.
To accelerate biological research with light microscopy, the Molecular BioEngineering Laboratory led by Kiryl D. Piatkevich at Westlake University is focused on the engineering, characterization, and validation of novel fluorescent proteins for advanced imaging applications. Since its establishment in 2019, the laboratory has already developed and reported several new fluorescent proteins, including the smallest green fluorescent proteins, miniGFPs1, a pair of bright blue fluorescent proteins, Electras2, and a set of sensitive fluorescent biosensors for recording potassium transient in mammalian cells3. More recently, the team led by Kiryl Piatkevich presented a systematic benchmarking of NIR FPs in various expression systems to aid end users in selecting the appropriate FPs for a given application4, which was published as a Registered Report by Nature Methods in 2023 (read “Behind the Paper” story here https://communities.springernature.com/posts/on-the-quest-for-the-fluorescent-superheroes).
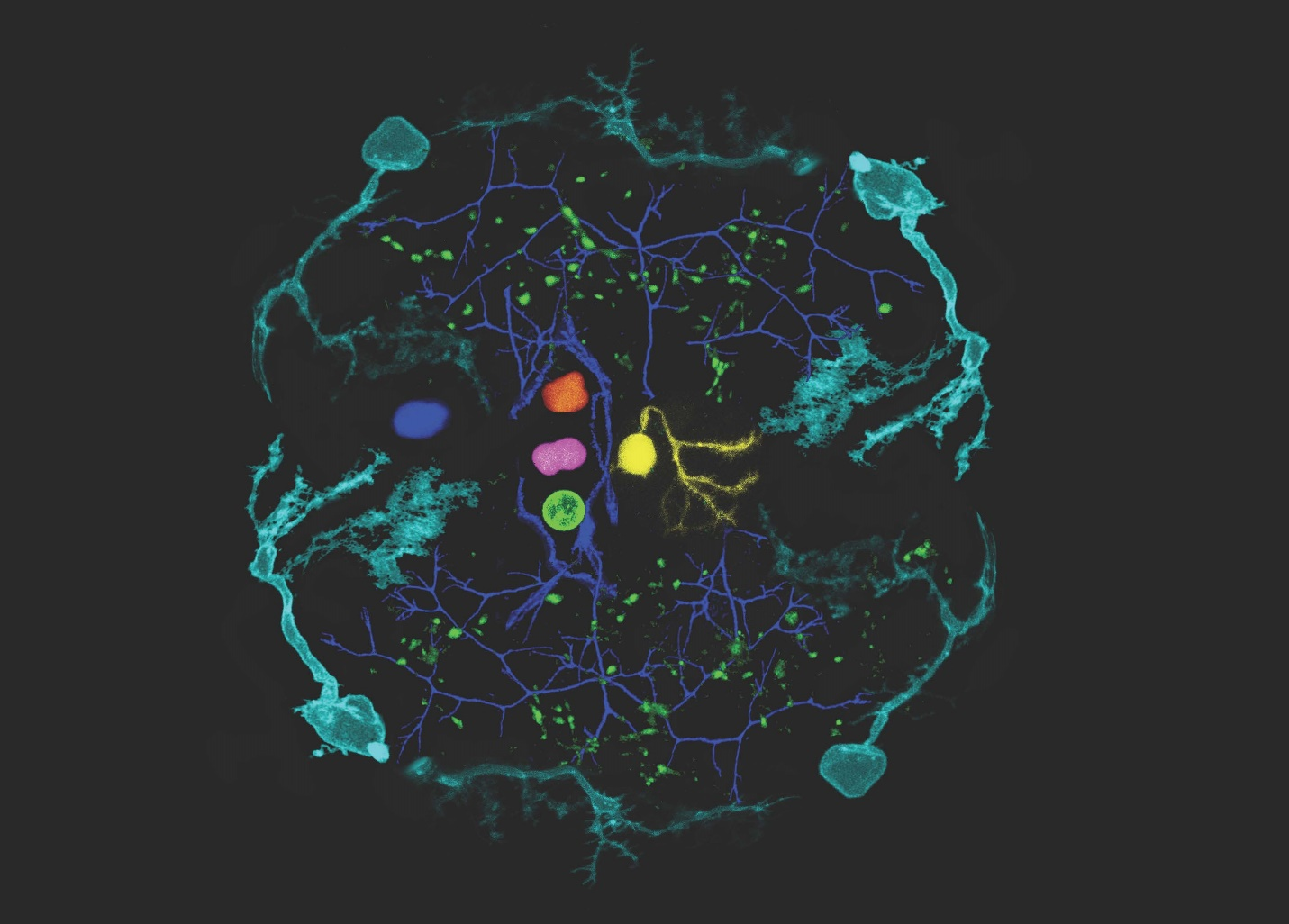
The edited fluorescence images of the zebrafish larval brain expressing achromatic variants of fluorescent proteins. The brain-shaped maze is surrounded by cerebellar Purkinje cells (Cyan) from zebrafish larvae, whereas the inner walls, which mimic cerebrovasculature, are made of long, growing axons of their motoneurons (Blue). The idea of this montage comes from 1980s video games with iconic neon-colored components. The major characters in this ‘game’ are made of cell bodies of hindbrain neurons (Green, Pink, Orange) showing beautiful, peculiar morphology, scoring points scattered in the maze, which are made of pre-synaptic signals in Purkinje cell axons (green puncta).(credits to Kazuhiko Namikawa from TU Braunschweig)
Kiryl Piatkevich says: “As a kid, I often argued with my friends about which superhero was the strongest and why. When I grew up, I replaced superheroes with FPs, and now I like to spend time on https://www.fpbase.org/ comparing different FPs (946 proteins to date) and helping my colleagues pick the right FP for various applications.” Similarly to the superheroes, diverse existing FPs have a variety of “superpowers” that can be utilized by biologists and microscopists to reveal the inner mechanisms of life. For example, flavin- and bilin-binding FPs can be expressed under anaerobic conditions, allowing live cell imaging of obligate anaerobic bacteria.1,5 The large Stokes shift red FPs are characterized by the highest pH stability and are thus beneficial for imaging in an acidic environment occurring in lysosomes and mitochondria.6–8

Fluorescent “superheroes”
However, among all the “superpowers” of fluorescent proteins, microscopists and cell biologists often opt for high intracellular brightness and photostability, which help gain better and longer visualization of target molecules and structures in cells. Therefore, for the past 30 years, protein engineers have been working hard to evolve new fluorescent proteins with enhanced brightness and photostability but have often been left disappointed at the trade-off between brightness and photostability: improved photostability results in lower brightness and vice versa. Serendipitously, naturally occurring green fluorescent protein from C. uchidae polyps exhibits exceptional photostability of an order of magnitude higher than that for any currently available FP while characterized by high molecular brightness, a unique combination in the world of fluorescent proteins. However, there is one obstacle to its application as a fluorescent tag for protein and membrane labeling: the discovered protein forms an obligate dimer, which can cause mislocalization and aggregation of fusions. To fix this problem, the research team led by Fedor Subach and Kiryl Piatkevich employed directed molecular evolution to disrupt the dimer interface in the target protein while preserving all other beneficial properties. The researchers succeeded in identifying a highly monomeric variant, which they called mBaoJin (‘bao jin’ means ‘stay gold’ in Mandarin). Compared to all other established and popular GFPs used for cell imaging, mBaoJin has 10-100 times higher photostability while exhibiting similar or higher brightness. This combination allows for boosting photon budget under imaging conditions and, as a result, improves the spatial and temporal resolution of observation. To realize this potential, the researchers performed long-term super-resolution imaging of various cellular structures using different super-resolution imaging modalities. It was possible to image α-tubulin and β-actin dynamics for up to 60 min without photobleaching using super-resolution microscopes – unprecedented performance for GFPs.
Movie - Long-term super-resolution imaging of actin dynamics with mBaoJin in HeLa cells using HIS-SIM imaging setup
Furthermore, the researchers found that the mBaoJin fluorescence resisted harsh chemical treatment with concentrate guanidinium chloride (one of the strongest denaturants used in physiochemical studies of protein folding). Extreme chemical stability is particularly beneficial for expansion microscopy (ExM), a super-resolution imaging technique where improvement in spatial resolution is achieved via physical sample magnification by infused swellable chemical polymer network9. Indeed, mBaoJin appeared to retain its high photostability and brightness in expanded samples, making it a protein of choice for ExM applications. Besides new advanced applications, mBaoJin opens an exciting opportunity to derive other spectral and photochemical phenotypes with enhanced photostability. To promote this direction, the researcher reported the high-resolution crystal structure of mBaoJin, which will inform the rational design of new derivatives. Additionally, mBaoJin may replace GFP-based reporting moiety in numerous genetically encoded green biosensors, enabling the development of next-generation photostable biosensors.
“We hope that mBaoJin will open a new chapter in fluorescent protein field initiating both advanced imaging applications and further engineering of deverse fluorescent protiens and biosensors” Kiryl Piatkevich adds.
Meanwhile, the research team embarks on a new adventure to find more fluorescent “superheroes” with blue, cyan, green, yellow, and red capes.
All plasmids reported in the publication are available from the WeKwikGene platform (https://wekwikgene.wllsb.edu.cn/) generously supported by Westlake Laboratory. WeKwikGene is the first plasmid repository in China. It was initiated by Prof. Piatkevich and funded by Westlake Laboratory of Life Sciences and Biomedicine. The available reagents are thoroughly tested and go through stringent quality control. On the website, you can find fully annotated, complete sequences for every plasmid. It offers a user-friendly experience, supports user account creation and Lab page, and provides the opportunity to increase the visibility of scientific work. WeKwikGene guarantees fast and free plasmid delivery of high-quality plasmids to support and promote open science.
References:
1. Liang, G.-T. et al. Enhanced small green fluorescent proteins as a multisensing platform for biosensor development. Front. Bioeng. Biotechnol. 10, 1039317 (2022).
2. Papadaki, S. et al. Dual-expression system for blue fluorescent protein optimization. Sci. Reports 2022 121 12, 1–16 (2022).
3. Torres Cabán, C. C. et al. Tuning the Sensitivity of Genetically Encoded Fluorescent Potassium Indicators through Structure-Guided and Genome Mining Strategies. ACS Sensors 7, 1336–1346 (2022).
4. Zhang, H. et al. Quantitative assessment of near-infrared fluorescent proteins. Nat. Methods 20, 1605–1616 (2023).
5. Chia, H. E., Zuo, T., Koropatkin, N. M., Marsh, E. N. G. & Biteen, J. S. Imaging living obligate anaerobic bacteria with bilin-binding fluorescent proteins. Curr. Res. Microb. Sci. 1, 1–6 (2020).
6. Erdogan, M., Fabritius, A., Basquin, J. & Griesbeck, O. Targeted In Situ Protein Diversification and Intra-organelle Validation in Mammalian Cells. Cell Chem. Biol. 27, 610-621.e5 (2020).
7. Subach, O. M. et al. LSSmScarlet, dCyRFP2s, dCyOFP2s and CRISPRed2s, Genetically Encoded Red Fluorescent Proteins with a Large Stokes Shift. Int. J. Mol. Sci. 2021, Vol. 22, Page 12887 22, 12887 (2021).
8. Vlaskina, A. M. ; V. ; et al. LSSmScarlet2 and LSSmScarlet3, Chemically Stable Genetically Encoded Red Fluorescent Proteins with a Large Stokes’ Shift. Int. J. Mol. Sci. 2022, Vol. 23, Page 11051 23, 11051 (2022).
9. Tillberg, P. W. et al. Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nat. Biotechnol. 34, 987–992 (2016).


















