



Search websites, locations, and people

Programmable gene therapies enabled by a novel translational regulation strategy
Mingqi Xie Lab
04, 2024
Email: zhangchi@westlake.edu.cn
Phone: +86-(0)571-86886861
Office of Public Affairs
The research group led by Dr. Mingqi Xie at Westlake University (MingLab) recently published a research article entitled “Engineered poly(A)-surrogates for translational regulation and therapeutic biocomputation in mammalian cells” in Cell Research. This study developed a new post-transcriptional gene regulation strategy that enables programmable control over translational initiation in human cells, substantially broadening the current repertoire and design principles of trigger-inducible gene switches and genetic sensors, hence taking a remarkable leap towards the future development of smartened gene therapies.

The paper can be read in full at https://doi.org/10.1038/s41422-023-00896-y
Almost all functions of our human body arise from a precise regulation of gene expression in living cells, where genetic information is programmed to pass through various control stages comprising epigenetic, transcriptional, translational or protein-level modifications. Gene regulation ensures that each cell expresses appropriate genes under appropriate developmental stages and under specific environmental conditions, thus maintaining the normal function of life. Synthetic biology adopts engineering principles to biological systems and seeks to regulate human cell functions through man-made control strategies of cellular gene expression. This lays the foundation for various gene therapy concepts, which use such engineered gene expression systems to treat certain genetic or metabolic disorders. At present, most gene regulation systems with such therapeutic utility act at the transcriptional level, which not only have relatively slow sense-and-response dynamics but are also generally limited to sense cellular events that occur within the nucleus. Post-transcriptional regulation systems, especially those operating at the level of protein translation, would be advantageous in many aspects, but the engineering of regulated translational devices with good therapeutic efficacy in vivo has long remained a critical hurdle in the field.
In this article, a research team led by Dr. Jiawei Shao, Shichao Li and colleagues developed a gene regulation strategy that permits programmable control over eukaryotic translational initiation. By excising the natural poly(A) signal of target genes and replacing it with a synthetic control region harboring RNA-binding protein (RBP)-specific aptamers, cap-dependent translation was rendered exclusively dependent on synthetic translation initiation factors containing different RBPs engineered to conditionally associate with different eIF4F-binding proteins. This not only marks a new regulation concept ensuring programmable, broadly adjustable and reversible (trans)gene expression control in mammalian cells with high efficacy and customizable durability in mice, but opened the door to a comprehensive design strategy for systematic engineering of gene switches and intracellular sensors with potential therapeutic utility. For example, placing insulin translation under the control of the FDA-approved drug grazoprevir resulted in effective control of blood glucose homeostasis in mice, demonstrating application potential for gene therapies that require on-demand drug action. For diseases that require instantaneous sense-and-response treatment programs, the translation-based regulation system was elegantly repurposed to create genetic sensors for various subcellularly (mis)localized protein markers (e.g. cancer-related fusion proteins), which could be either used alone or in combination with other cell-state classification strategies to create therapeutic biocomputers to drive self-sufficient elimination of tumor cells. Taken together, this work describes an undiscovered strategy for translational regulation, adds intracellular protein sensing to the portfolio of cell-state classification strategies and could accelerate the clinical translation of various programmable gene therapies.
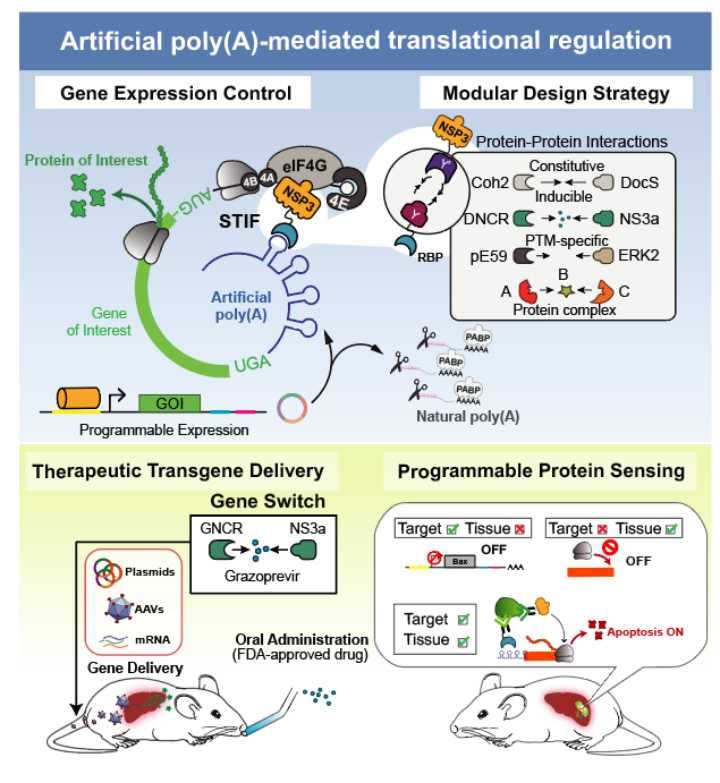
Fig. 1 shows a schematic illustration of the synthetic translational regulation system construction and application. To achieve the concept of “controllable circularization” in translation regulation, the researchers designed a synthetic translation initiation system composed of custom-designed mRNA and synthetic translation initiation factors (STIFs). The custom-designed mRNA contains RNA-binding protein (RBP)-specific aptamers in the 3’-untranslated region (3’ UTR), followed by a ribonuclease sequence or a shRNA target site to remove the poly-A tail from the mRNA to insulate it from endogenous translation processes. STIFs consist of RBPs, specifically binding with 3’-UTR, and rotaviral non-structural protein 3 (NSP3), interacting with eIF4G with high affinity. The conditionally assemble of the two independent proteins of STIFs via protein-protein interactions can trigger the circularization of target mRNA for translation initiation. The modular design framework enables systematic engineering of trigger inducible gene switches and programmable protein sensors that respond to various intracellular biomarkers. Researchers successfully constructed a translation-based gene switch triggered by the FDA-approved drug grazoprevir which can effectively control insulin expression and restore glucose homeostasis in diabetic mice. Researchers also constructed a translation-based protein sensor and combined it with tissue specific promoter to create therapeutic biocomputers driving self-sufficient elimination of tumor cells in mice.
RELATED
NEWS
Programmable gene therapies enabled by a novel translational regulation strategy


















